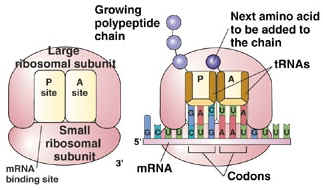
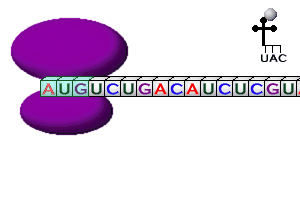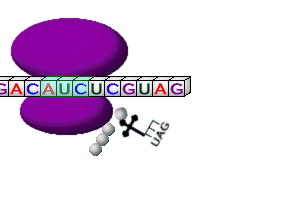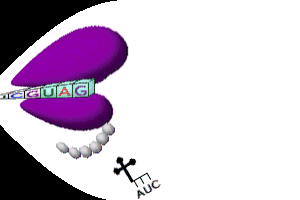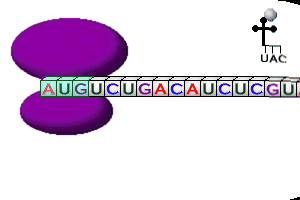
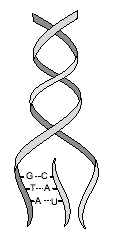The Nobel Prize in Physiology or Medicine for 2004 awarded
jointly to
Richard Axel and Linda B. Buck
for their discoveries of
“odorant receptors and the organization of the olfactory system”
Introduction
The sense of smell long remained the most enigmatic of our senses. The basic principles for recognizing and remembering about 10,000 different odors were not understood. This year’s Nobel Laureates in Physiology or Medicine have solved this problem and in a series of pioneering studies clarified how our olfactory system works. They discovered a large gene family, comprised of some 1,000 different genes (three per cent of our genes) that give rise to an equivalent number of olfactory receptor types. These receptors are located on the olfactory receptor cells, which occupy a small area in the upper part of the nasal epithelium and detect the inhaled odorant molecules.
Each olfactory receptor cell possesses only one type of odorant receptor, and each receptor can detect a limited number of odorant substances. Our olfactory receptor cells are therefore highly specialized for a few odors. The cells send thin nerve processes directly to distinct micro domains, glomeruli, in the olfactory bulb, the primary olfactory area of the brain. Receptor cells carrying the same type of receptor send their nerve processes to the same glomerulus. From these micro domains in the olfactory bulb the information is relayed further to other parts of the brain, where the information from several olfactory receptors is combined, forming a pattern. Therefore, we can consciously experience the smell of a lilac flower in the spring and recall this olfactory memory at other times.
Richard Axel, New York, USA, and Linda Buck, Seattle, USA, published the fundamental paper jointly in 1991, in which they described the very large family of about one thousand genes for odorant receptors. Axel and Buck have since worked independent of each other, and they have in several elegant, often parallel, studies clarified the olfactory system, from the molecular level to the organization of the cells.
The olfactory system is important for life quality
When something tastes really good it is primarily activation of the olfactory system which helps us detect the qualities we regard as positive. A good wine or a sun ripe wild strawberry activates a whole array of odorant receptors, helping us to perceive the different odorant molecules.
A unique odor can trigger distinct memories from our childhood or from emotional moments – positive or negative – later in life. A single clam that is not fresh and will cause malaise can leave a memory that stays with us for years, and prevent us from ingesting any dish, however delicious, with clams in it. To lose the sense of smell is a serious handicap – we no longer perceive the different qualities of food and we cannot detect warning signals, for example smoke from a fire.
Olfaction is of central importance for most species
All living organisms can detect and identify chemical substances in their environment. It is obviously of great survival value to be able to identify suitable food and to avoid putrid or unfit foodstuff. Whereas fish has a relatively small number of odorant receptors, about one hundred, mice – the species Axel and Buck studied – have about one thousand. Humans have a somewhat smaller number than mice; some of the genes have been lost during evolution.
Smell is absolutely essential for a newborn mammalian pup to find the teats of its mother and obtain milk – without olfaction the pup does not survive unaided. Olfaction is also of paramount importance for many adult animals, since they observe and interpret their environment largely by sensing smell. For example, the area of the olfactory epithelium in dogs is some forty times larger than in humans.
A large family of odorant receptors
The olfactory system is the first of our sensory systems that has been deciphered primarily using molecular techniques. Axel and Buck showed that three per cent of our genes are used to code for the different odorant receptors on the membrane of the olfactory receptor cells. When an odorant receptor is activated by an odorous substance, an electric signal is triggered in the olfactory receptor cell and sent to the brain via nerve processes. Each odorant receptor first activates a G protein, to which it is coupled. The G protein in turn stimulates the formation of cAMP (cyclic AMP). This messenger molecule activates ion channels, which are opened and the cell is activated. Axel and Buck showed that the large family of odorant receptors belongs to the G protein-coupled receptors (GPCR).
All the odorant receptors are related proteins but differ in certain details, explaining why they are triggered by different odorous molecules. Each receptor consists of a chain of amino acids that is anchored into the cell membrane and traverses it seven times. The chain creates a binding pocket where the odorant can attach. When that happens, the shape of the receptor protein is altered, leading to G protein activation.
One type of odorant receptor in each olfactory receptor cell
Independently, Axel and Buck showed that every single olfactory receptor cell expresses one and only one of the odorant receptor genes. Thus, there are as many types of olfactory receptor cells as there are odorant receptors. It was possible to show, by registering the electrical signals coming from single olfactory receptor cells, that each cell does not react only to one odorous substance, but to several related molecules – albeit with varying intensity.
Buck’s research group examined the sensitivity of individual olfactory receptor cells to specific odorants. By means of a pipette, they emptied the contents of each cell and showed exactly which odorant receptor gene was expressed in that cell. In this way, they could correlate the response to a specific odorant with the particular type of receptor carried by that cell.
Most odors are composed of multiple odorant molecules, and each odorant molecule activates several odorant receptors. This leads to a combinatorial code forming an “odorant pattern” – somewhat like the colors in a patchwork quilt or in a mosaic. This is the basis for our ability to recognize and form memories of approximately 10,000 different odors.
Olfactory receptor cells activate micro regions in the olfactory bulb
The finding that each olfactory receptor cell only expresses one single odorant receptor gene was highly unexpected. Axel and Buck continued by determining the organization of the first relay station in the brain. The olfactory receptor cell sends its nerve processes to the olfactory bulb, where there are some 2,000 well-defined microregions, glomeruli. There are thus about twice as many glomeruli as the types of olfactory receptor cells.
Axel and Buck independently showed that receptor cells carrying the same type of receptor converge their processes into the same glomerulus, and Axel’s research group used sophisticated genetic technology to demonstrate in mice the role of the receptor in this process. The convergence of information from cells with the same receptor into the same glomerulus demonstrated that also glomeruli exhibit remarkable specificity (see figure).
In the glomeruli we find not only the nerve processes from the olfactory receptor cells but also their contacts with the next level of nerve cells, the mitral cells. Each mitral cell is activated only by one glomerulus, and the specificity in the information flow is thereby maintained. Via long nerve processes, the mitral cells send the information to several parts of the brain. Buck showed that these nerve signals in turn reach defined micro regions in the brain cortex. Here the information from several types of odorant receptors is combined into a pattern characteristic for each odor. This is interpreted and leads to the conscious experience of a recognizable odor.
Pheromones and taste
The general principles that Axel and Buck discovered for the olfactory system appears to apply also to other sensory systems. Pheromones are molecules that can influence different social behaviors, especially in animals. Axel and Buck, independent of each other, discovered that pheromones are detected by two other families of GPCR, localized to a different part of the nasal epithelium. The taste buds of the tongue have yet another family of GPCR, which is associated with the sense of taste.
Reference: Buck, L. and Axel, R. (1991) Cell, vol. 65, 175-187.
Odorant Receptors and the Organization of the Olfactory System
Source: http://nobelprize.org/nobel_prizes/medicine/laureates/2004/press.html